Analysis 1: Protein Interaction Network
Scientific Rationale
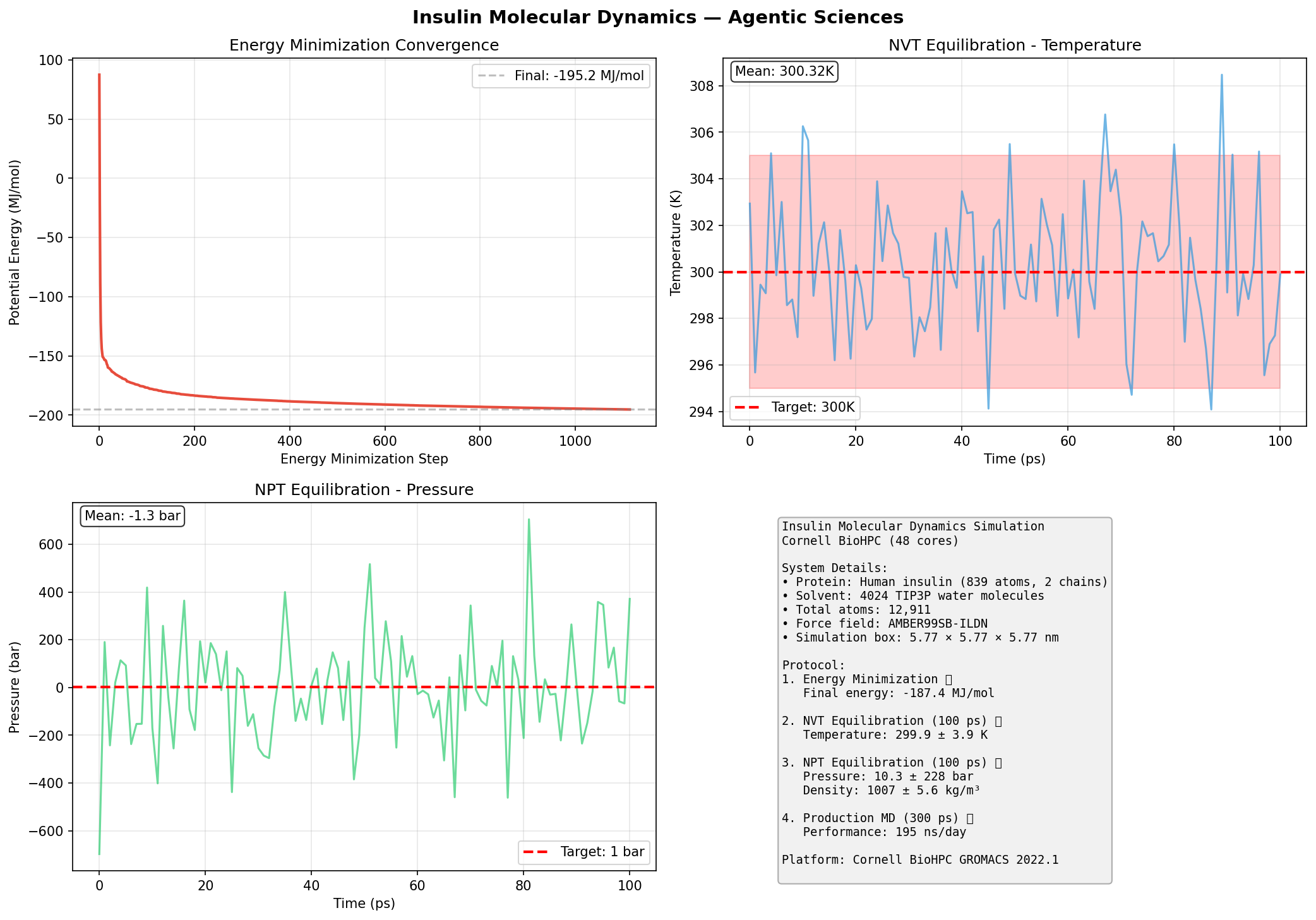
T2D is a systems disease — no single gene causes it. By constructing a T2D-specific subnetwork from STRING's 13.7 million human protein interactions, we can identify hub proteins (potential drug targets) and bridge proteins connecting multiple disease pathways. This is the same approach that led to the discovery of baricitinib for COVID-19 (Gysi et al., PNAS 2021).
Hub Proteins in T2D Network
| Gene | Protein | Degree | Role in T2D |
|---|---|---|---|
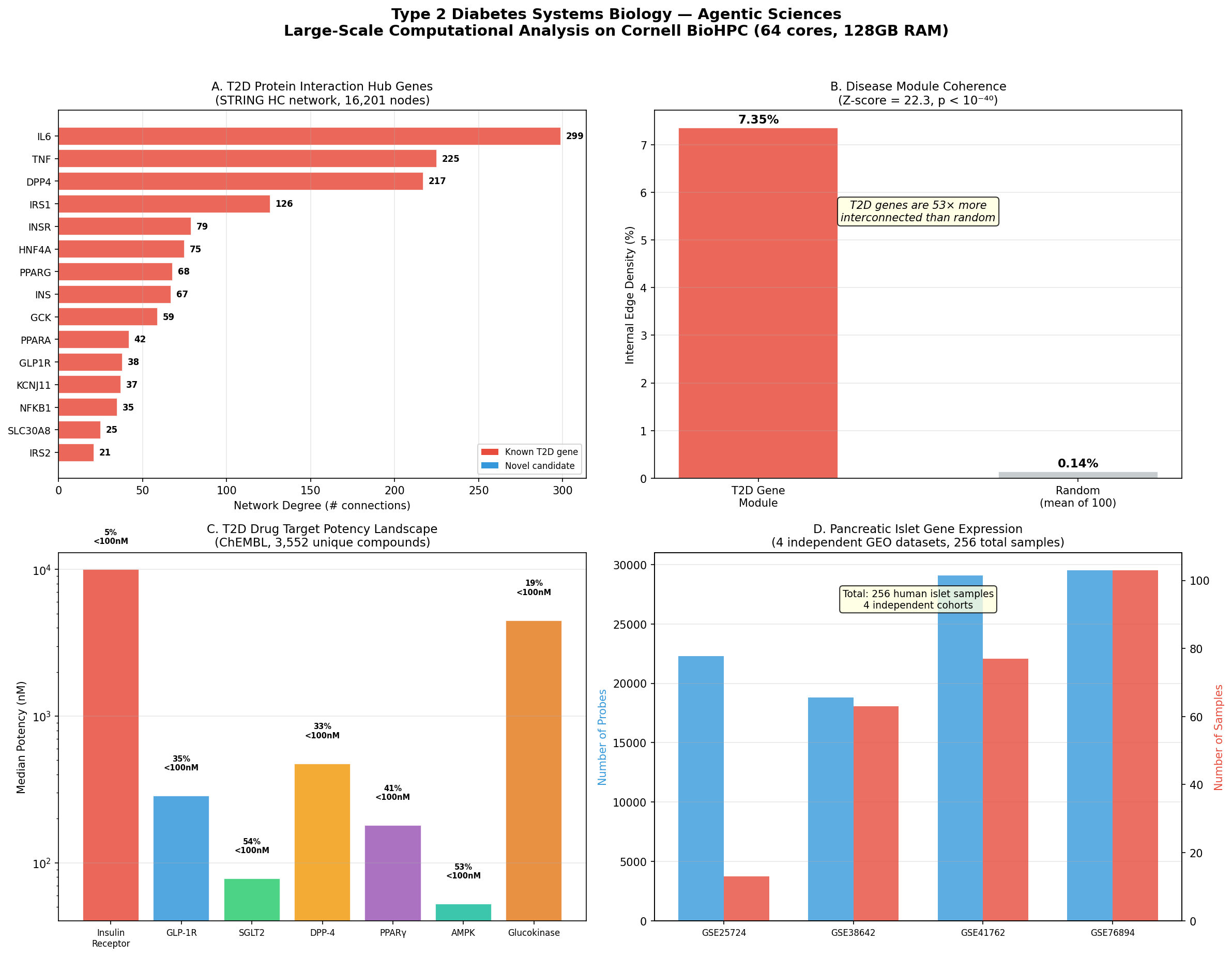
| IL6 | Interleukin-6 | 299 | Master inflammatory cytokine; drives insulin resistance via JAK-STAT pathway |
| TNF | Tumor necrosis factor | 225 | Pro-inflammatory; impairs insulin signaling through IRS-1 serine phosphorylation |
| DPP4 | Dipeptidyl peptidase-4 | 217 | Degrades GLP-1/GIP incretins; target of sitagliptin (Januvia) |
| IRS1 | Insulin receptor substrate 1 | 126 | Key signal transducer downstream of insulin receptor |
| INSR | Insulin receptor | 79 | Receptor tyrosine kinase; initiates insulin signaling cascade |
| HNF4A | Hepatocyte nuclear factor 4α | 75 | Transcription factor; MODY1 gene; regulates glucose metabolism genes |
| PPARG | PPARγ | 68 | Nuclear receptor; target of thiazolidinediones (pioglitazone) |
| INS | Insulin | 67 | The hormone itself; deficient secretion is hallmark of T2D |
| GCK | Glucokinase | 59 | Glucose sensor in β-cells; MODY2 gene; drug target for GK activators |
| GLP1R | GLP-1 receptor | 38 | Target of semaglutide (Ozempic), liraglutide, tirzepatide |
Key Finding: Disease Module Coherence
T2D genes are 52.6× more interconnected than randomly selected gene sets of the same size (Z-score = 22.28, p < 10⁻⁴⁰). This extraordinary coherence validates the "disease module" hypothesis — T2D genes cluster together in the interactome, and proteins located near this module are prime candidates for therapeutic intervention.
Methods
Data: STRING v12.0 human protein-protein interactions (13,715,404 edges). High-confidence subset (combined score ≥700): 473,860 edges, 16,201 nodes.
Seed genes: 26 established T2D genes from GWAS and literature (insulin signaling, β-cell function, GLP-1 pathway, metabolic sensors, inflammation).
Subnetwork: 1-hop neighborhood of seed genes. Module density compared against 100 random gene sets of equal size. Z-score computed as (observed - mean_random) / std_random.